Scientific Computing and Data / High Performance Computing / Hardware and Technical Specs
Hardware and Technical Specs
- The Minerva supercomputer is maintained by Scientific Computing and Data (SCD) at the Icahn School of Medicine, Mount Sinai.
- Minerva was created in 2012 and has been upgraded several times (most recently in Nov. 2024 and Feb. 2026) and has over 20 petaflops of compute power.
- It consists of 25,584 Intel Platinum processors in different generations including 2.1 GHz, 2.3GHz, 2.6 GHz, and 2.9 GHz compute cores (112 cores or 96 cores or 64 cores or 48 cores per node with two sockets in each node) with 1.5 terabytes (TB) or 2 TB of memory per node. In addition, Minerva also has 408 graphics processing units (GPUs), including 48 Nvidia B200s, 236 H100 Nvidia GPUs, 32 L40S Nvidia servers, 44 A100 Nvidia GPUs, 48 V100 Nvidia GPUs, 452 TB of total memory, and 32 petabytes of spinning storage accessed via IBM’s Spectrum Scale/General Parallel File System (GPFS).
- Minerva has contributed to over 2,100 peer-reviewed publications since 2012.
The following diagram shows the overall Minerva configuration:
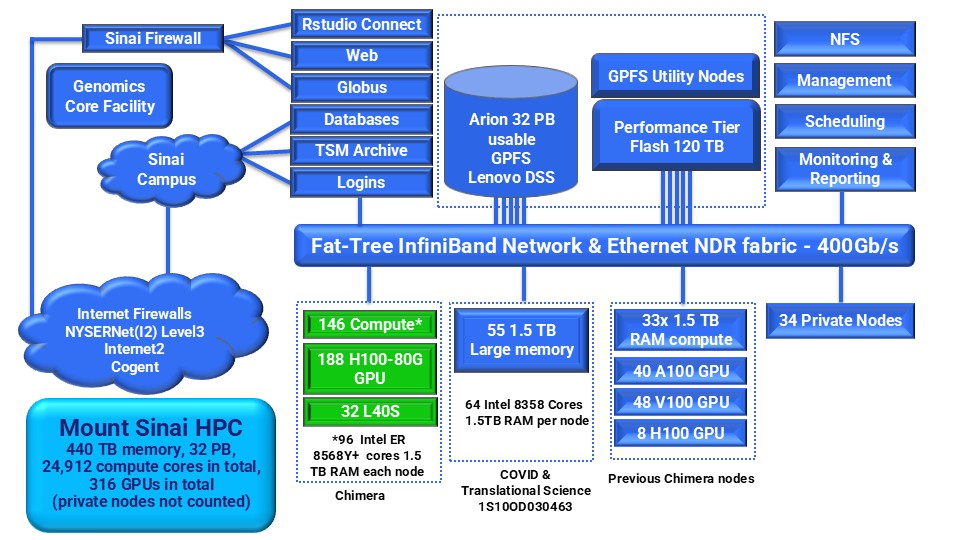
Compute Nodes
Chimera Partition
| Added in Nov. 2024. | Nodes purchased prior to 2024 and integrated to new NDR network via HDR 100Gb/s. |
|
|
| Added in Feb. 2026. | |
|
BODE2 Partition |
CATS Partition |
AIMS Partition |
[Decommissioned on July 17th and Nov. 5th, 2024].$2M S10 BODE2 awarded by NIH (Kovatch PI).
|
$2M CATS awarded by NIH (Kovatch PI).
|
$2M AIMS awarded by NIH (Kovatch PI), launched in Feb, 2026
|
Summary
| Total system memory (computes + GPU) = 452 TB | Total number of cores (computes + GPU) = 25,584 cores |
| CPU Peak performance of all nodes = > 1.9 PFLOPS | H100 Peak performance based FP64 Tensor cores = 15.2 PFLOPS. |
File System Storage
Minerva uses IBM’s General Parallel File System (GPFS) because it has advantages that are specifically useful for informatics workflows that involve high speed metadata access, tiered storage, and sub-block allocation. Metadata is the information about the data in the file system, and it is stored in flash memory for fast access. A parallel file system was used for Minerva because NFS and other file systems cannot scale to the number of nodes or provide performance for the large number of files involved in typical genomics workflows.
Currently we have one parallel file system on Minerva, Arion, which users can access at /sc/arion. The Hydra file system was retired at the end of 2020.
| GPFS Name | Lifetime | Storage Type | Raw PB | Usable PB |
| Arion | 2019 – | Lenovo DSS | 14 | 9.6 |
| Arion | 2019 – | Lenovo G201 flash | 0.12 | 0.12 |
| Arion | 2020 – | Lenovo DSS | 16 | 11.2 |
| Arion | 2021 – | Lenovo DSS | 16 | 11.2 |
| Total | 46 | 32 |
Acknowledging Mount Sinai in Your Work
This work was supported by grant UL1TR004419 from the National Center for Advancing Translational Sciences, National Institutes of Health.
Using the S10 BODE and CATS Minerva partitions requires acknowledgements of support by NIH in your publications. To assist, we have provided exact wording of acknowledgements required by NIH for use in publications and other work. Click here to learn how to acknowledge Minerva and NIH support in your publications.
