TCGA – The Cancer Genome Atlas Program
Overview
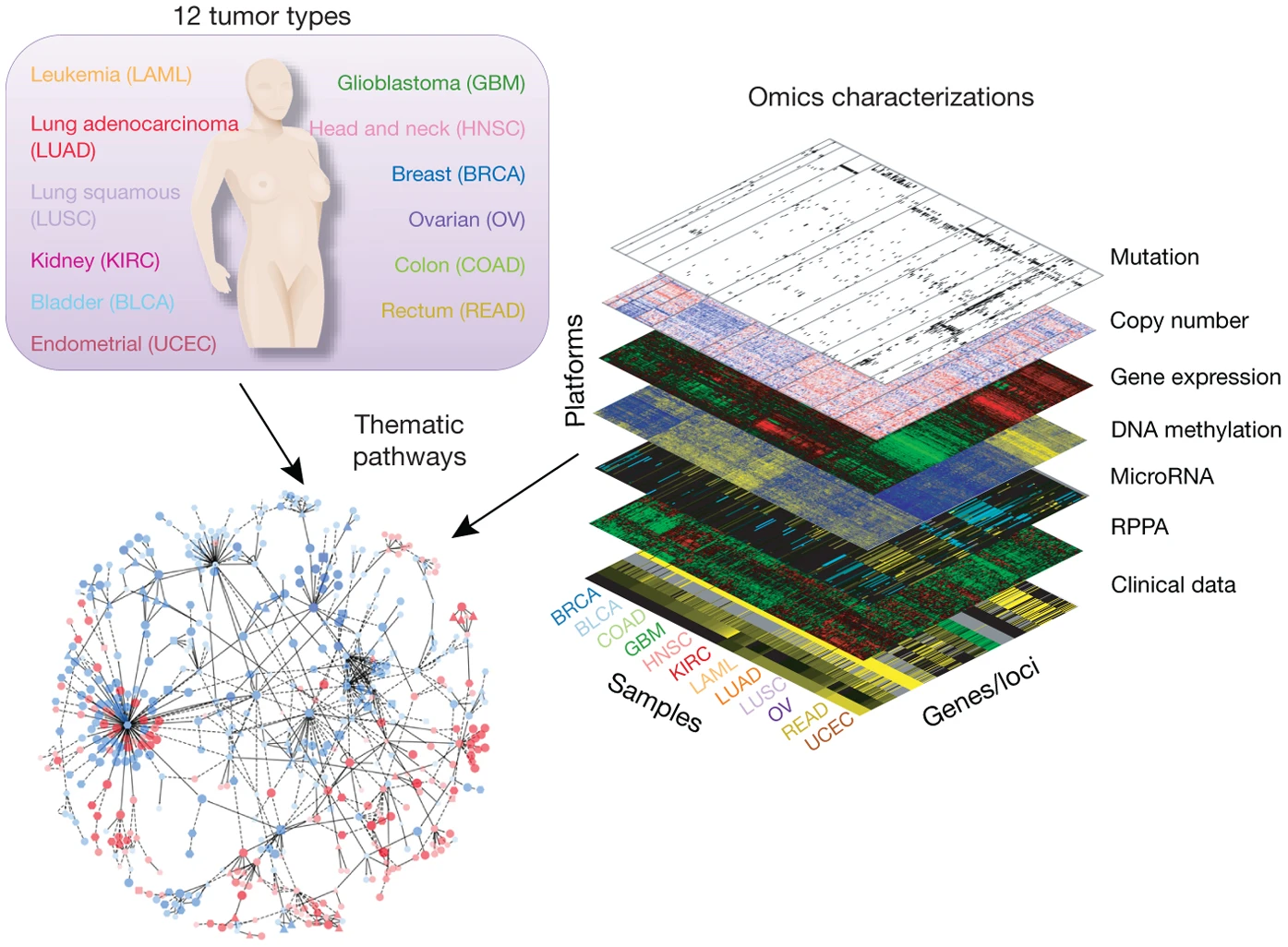
The Cancer Genome Atlas (TCGA) is a landmark cancer genomics program that molecularly characterized over 20,000 primary cancer and matched normal samples spanning 33 cancer types. The program is a joint effort between National Cancer Institute and the National Human Genome Research Institute first established in 2016.
Currently, two versions are hosted on the Data Ark: Version 31.0 and version 32.0. The gene model used as a reference across TCGA has been updated from GENCODE 22(GRC37/hg19)—version 31 to GENCODE 36 (GRCh38/hg38)–version 32. To learn more about the data sets from a different version, find the data release notes here.
All the TCGA data sets belong to the “open-access” category and were obtained from the Genomic Data Commons Data Portal. The TCGA folder on Minerva Supercomputer hosts all the biospecimen, clinical, RNA-seq counts, WXS -Mutation Annotation Format (MAF), and the TCGA data sets from cBioPortal.
TCGA Processed Data Sets
TCGA data set has been processed and uploaded to Data Ark by Dr.Deniz Demircioglu (deniz.demircioglu@mssm.edu), who combined data from the biospecimen and clinical folder and consolidated all the RNA-seq counts files (over 11,000 patients) into 33 different outcomes. See the table below. For more information about the TCGA Study Abbreviations, click here.
| Abbrev. | Study Name | Abbrev. | Study Name | Abbrev. | Study Name |
| ACC | Adrenocortical carcinoma | BLCA | Bladder Urothelial carcinoma | BRCA | Breast invasive carcinoma |
| CESC | Cervical squamous cell carcinoma and endocervical adenocarcinoma | CHOL | Cholangiocarcinoma | COAD | Colon adenocarcinoma |
| DLBC | Lymphoid Neoplasm Diffuse Large B-cell Lymphoma | ESCA | Esophageal carcinoma | GBM | Glioblastoma multiforme |
| HNSC | Head and Neck squamous cell carcinoma | KICH | Kidney Chromophobe | KIRC | Kidney renal clear cell carcinoma |
| KIRP | Kidney renal papillary cell carcinoma | LAML | Acute Myeloid Leukemia | LGG | Brain Lower Grade Glioma |
| LIHC | Liver hepatocellular carcinoma | LUAD | Lung adenocarcinoma | LUSC | Lung squamous cell carcinoma |
| MESO | Mesothelioma | OV | Ovarian serous cystadenocarcinoma | PAAD | Pancreatic adenocarcinoma |
| PCPG | Pheochromocytoma and Paraganglioma | PRAD | Prostate adenocarcinoma | READ | Rectum adenocarcinoma |
| SARC | Sarcoma | SKCM | Skin Cutaneous Melanoma | STAD | Stomach adenocarcinoma |
| TGCT | Testicular Germ Cell Tumors | THCA | Thyroid carcinoma | THYM | Thymoma |
| UCEC | Uterine Corpus Endometrial Carcinoma | UCS | Uterine Carcinosarcoma | UVM | Uveal Melanoma |
Inside each outcome folder, you will see the following files:
| aliquot.tsv | count.tsv | fpkm.tsv | sample.tsv |
| analyte.tsv | exposure.tsv | fpkm_uq.tsv | slide.tsv |
| clinical.tsv | family_history.tsv | portion.tsv |
Access
Effective from January 22, 2024, you must read, agree and sign the Data Use Agreement (you must be logged in through the Mount Sinai campus network or secure remote VPN). Access is granted within 24 hours, and on Minerva, you can load module $ module load dataark to see the path variables.
By using these data, you agree to acknowledge BiNGS–the Tisch Cancer Institute Bioinformatics Core for Next-Generation Sequencing and the Data Ark team for all oral and written presentations, grand submission, awards, and publications resulting from any analyses of the data sets.
Data Ark Data Sets
Please visit the Data Ark Data Set webpage to explore other data sets.
