Scientific Computing and Data / Research Data Services / Data Ark: Data Commons / Mount Sinai COVID-19 EHR Data
Mount Sinai Data Warehouse COVID-19 Electronic Health Record (EHR) Data Set
Overview
To support COVID-related research at Mount Sinai utilizing electronic health record data, the Mount Sinai Data Warehouse team created this near real-time and evolving de-identified COVID-19 data set. The cohort includes patients with an encounter at a Mount Sinai facility who have been diagnosed with COVID-19, those who are under investigation for COVID-19, as well as those who have screened negative for COVID-19. The data set consists of multiple files (patient encounter, lab tests, medication administrations, diagnoses, vitals, radiology and immunizations) all of which are linked together using a masked medical record number and a unique patient encounter key. This data set is updated every day.
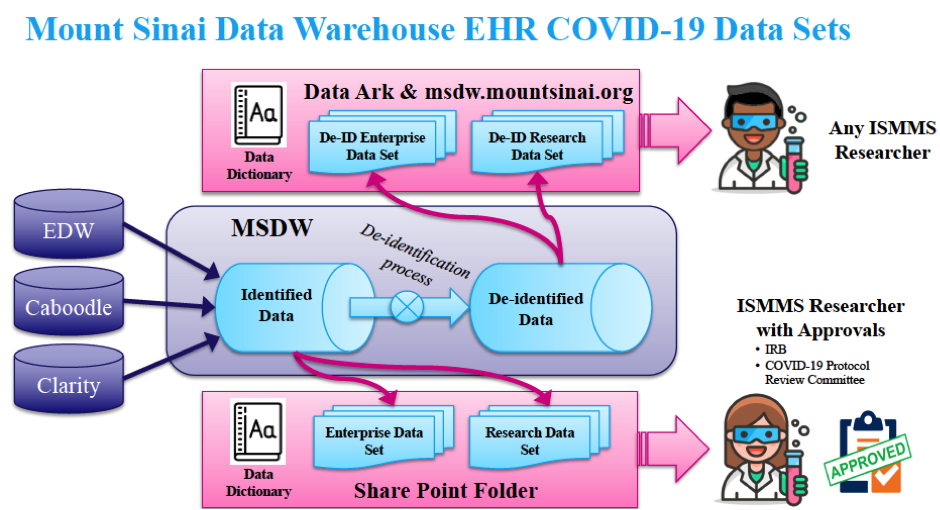
Data Sources and De-identification Process for HIPAA Privacy Rule Compliance
The data set is derived from Mount Sinai’s Caboodle, Clarity, and Enterprise Data Warehouse databases. To be transparent and to promote reproducibility and usability, a detailed data dictionary was created to accompany the data sets. The data dictionary contains information on the definition of each data element in the set, as well as a log of all changes made.
In the de-identified data set, all 18 protected health information identifiers as delineated in the HIPAA Privacy Rule, including names, dates and addresses, are masked or removed. In order to maintain the temporal relationship between events, all dates are converted to elapsed days relative to the COVID encounter start date.
Process for COVID-19 Data Sets
- Mask or remove all 18 PHI identifiers
- Birth date converted to age at encounter, with all ages over 89 set to 90
- Convert all dates to elapsed days relative to encounter date
- Zip codes truncated to 3 digits (or masked if population < 20k)
Process for Other MSDW Data Sets
- Same as above, but all dates for each patient are shifted by a random integer to preserve chronology

De-identified COVID-19 Research Data Set Elements and Format
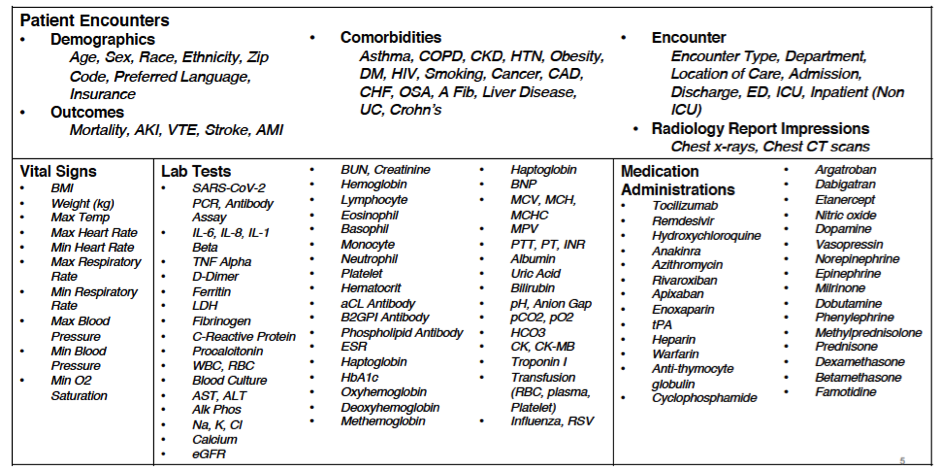
A COVID related visit is one in which the patient had a SARS-CoV-2 PCR test ordered or resulted, a diagnosis of COVID-19 or a COVID related reason for visit. As this data set was meant to be used for a wide variety of research purposes, the inclusion criteria cast a wide net and includes all patients associated with the pandemic and not just those who tested positive for COVID. Researchers can refine the cohort to exclude patients as needed. The data sets are structured so that new data elements, that fit into the existing data structure, can easily be added. For example, it is relatively easy to add additional lab results or medications, which has been important as evolving research and clinical experience highlight new therapeutics and biomarkers key to treating and researching COVID patients. Additionally, custom data sets can be created that include data elements that fall outside of the standard file structure, such as data set oxygen device files or free text clinical notes.
Value Sets for Easier Usage
Equivalent data element components are mapped to a single group as a value set. For instance, all of the neutrophil-related counts are included in the neutrophil # component type group.
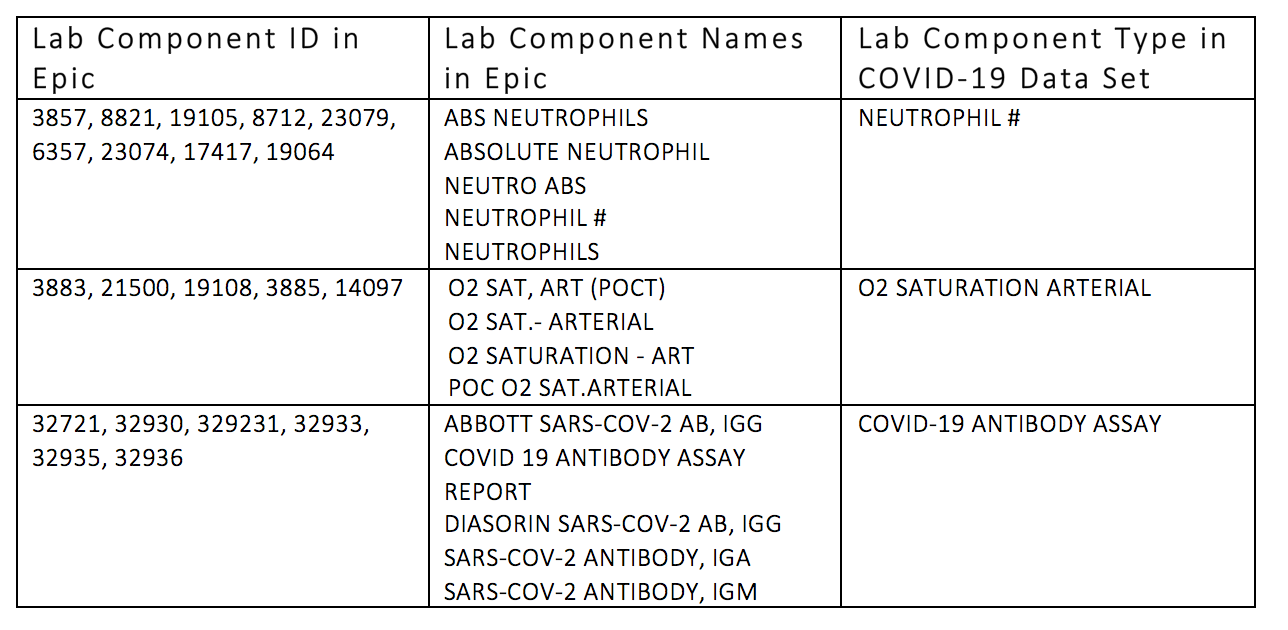
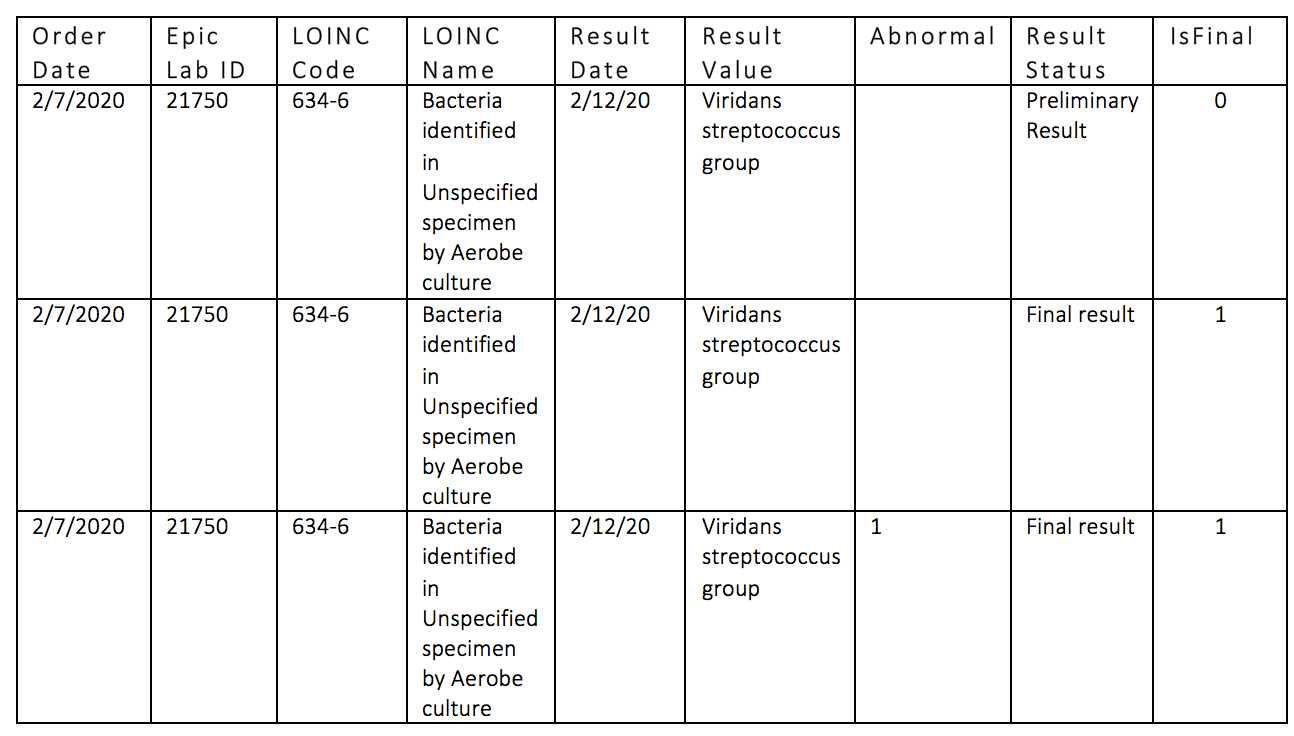
Record Updates Are Handled Over Time
As lab results are reported or updated, the newest result appears in the lab file, overwriting the older values.
Access
To access the identified version of the COVID data set, the user must have approval from the IRB and/or the appropriate QI board, as well as the internal Mount Sinai COVID-19 Protocol Review Committee. Once these documents and approvals are obtained, please open a data request attaching the appropriate IRB or QI board approval letter(s) at https://msdw.mountsinai.org/.
To use this data, you must read, agree and sign the Data Use Agreement (you must be logged in through the Mount Sinai campus network or secure remote VPN). On Minerva, you can load module $ module load dataark to see the path variables.
Data Ark Data Sets
Please visit the Data Ark Data Set webpage to explore other data sets.
