Scientific Computing and Data / Research Data Services / Data Ark: Data Commons / 1,000 Genomes Project
1,000 Genomes Project
Overview
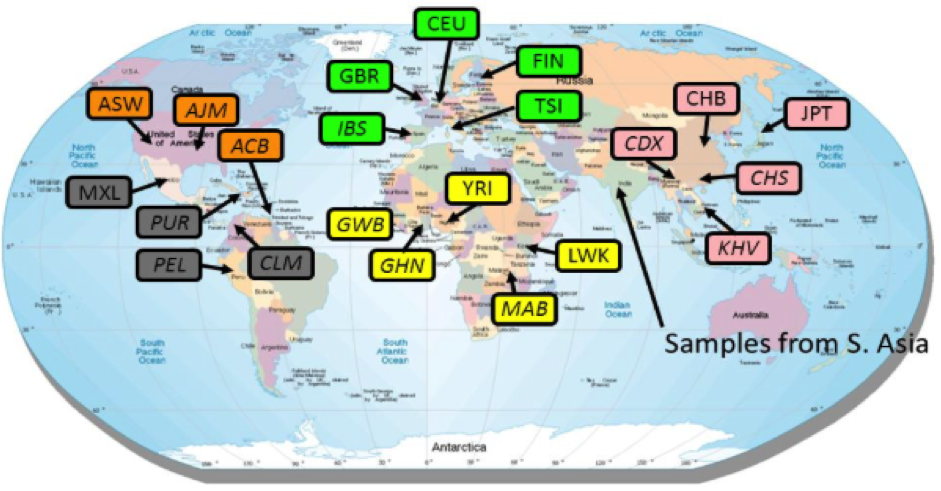
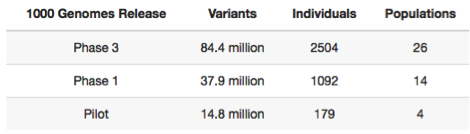
The 1000 Genomes Project was initiated in 2008, comprising 3 pilot phases that focused on low coverage whole genome sequencing (WGS) of 180 individuals of African/Asian/European ancestry, and deep coverage of two trios and of 1000 genes in 900 unrelated samples. These pilot studies were expanded to larger projects, published across three Nature papers published in 2010, 2012, 2015, with the final data set corresponding to 2,504 individuals from 26 global populations (~4X WGS and 30X WES for all, and 24 individuals with 30X WGS for validation).
Since its completion in 2013, the 1000 Genomes Project was superseded by The International Genome Sample Resource (IGSR), established to host and extend the 1000G data. For more information on the 1000G Project and the IGSR see their website here: 1000G & IGSR website.
The Data Ark hosts a replica of the Phase 3 individual-level called genotype data (VCF format) created by Google Health using high coverage (30X) Illumina sequencing performed by the New York Genome Center. The CRAM files can be obtained from the 1000G website but these are extremely large (36TB) and will not be needed for the vast majority of user cases and so are not included on the Data Ark.
Access
Effective from January 22, 2024, you must read, agree and sign the Data Use Agreement (you must be logged in through the Mount Sinai campus network or secure remote VPN). Access is granted within 24 hours, and on Minerva, you can load module $ module load dataark to see the path variables.
Data Ark Data Sets
Please visit the Data Ark Data Set webpage to explore other data sets.
