Over time, some of the Ma’ayan Lab’s tools have been retired as newer, more advanced platforms replaced them. While no longer maintained, these deprecated tools played an important role in shaping the current ecosystem. They introduced pioneering approaches in computational systems biology and bioinformatics.
Browse through the following categories to explore the various original resources we have developed:
Most Popular | Recently Published | Enrichment Analysis | Drug and Target Discovery | Computational Platforms and Workflow Engines | Gene and Drug Pages | lncRNAs | Data Portals | Data Visualization Components | Deprecated
LJP-BCNB
LINCS Joint Project – Breast Cancer Network Browser
LJP-BCNB visualizes thousands of signatures from six breast cancer cell lines treated with ~100 single molecule perturbations, mostly kinase inhibitors. These perturbations were applied in different concentrations while gene expression was measured at different time points using the L1000 technology. Under the same conditions, the cells were imaged for cell viability. The distance between nodes represents response similarity computed using the cosine distance between the Characteristic Direction vectors of perturbations compared with their appropriate controls.
PMID: 29084964
COVID-19 Drug and Gene Set Library
Collection of Drug and Gene Sets Related to COVID-19 Contributed by the Community
Enables users to view, download, analyze, visualize, and contribute drug and gene sets related to COVID-19 research. To evaluate the content of the library, we compared the results from six in vitro drug screens for COVID-19 repurposing candidates. Surprisingly, we observe low overlap across screens while highlighting overlapping candidates that should receive more attention as potential therapeutics for COVID-19. Overall, the COVID-19 Drug and Gene Set Library can be used to identify community consensus, make researchers and clinicians aware of new potential therapies, enable machine-learning applications, and facilitate the research community to work together toward a cure.
PMID: 32838343
Principal Angle Enrichment Analysis
Dimensionally Reduced Multivariate Gene Set Enrichment Analysis Tool
Uses the geometrical concept of the principal angle to quantify gene-set enrichment. We find that PAEA outperforms a selection of commonly used gene set enrichment methods including GSEA. To benchmark PAEA with other enrichment methods we use real data. We examined the ranking of transcription factors by performing enrichment analysis on gene expression signatures from many studies that knocked-down, knocked-out or over-expressed transcription factors, and performed the enrichment analysis with a library of gene sets created from ChIP-Seq data profiling the same transcription factors.
PMID: 26848405
Slicr
LINCS L1000 Slicr
A metadata search engine that searches for LINCS L1000 gene expression profiles and signatures matching users’ input parameters. It features download of selected search results as csv files in a zipped folder and visualization of selected results in a 3D scatter plot using PCA or MDS. Slicr consists of three views: the search view, the checkout view and the 3D scatter view.
LINCS Canvas Browser
LINCS L1000 Clustering, Visualization and Enrichment Analysis Tool
A web-based tool that enables users to explore thousands of genome-wide gene expression experiments applied to breast cancer cell lines. The browser visualizes results from L1000 experiments where drugs or endogenous ligands were applied to six human breast cancer cell lines in different concentrations and where expression was measured at different time points. The visualization of the results is organized by cell-line and batch where perturbations that induced similar responses are clustered together on a canvas.
PMID: 24906883
Drug/Cell-line Browser
Data Visualization Tool
An interactive HTML5 data visualization tool for interacting with three of the recently published datasets of cancer cell lines/drug-viability studies. DCB uses clustering and canvas visualization of the drugs and the cell lines, as well as a bar graph that summarizes drug effectiveness for the tissue of origin or the cancer subtypes for single or multiple drugs. DCB can help in understanding drug response patterns and prioritizing drug/cancer cell line interactions by tissue of origin or cancer subtype.
PMID: 25100688
Cite-D-Lite
Chrome Extension for Data and Paper Citations with Text Importance Highlighting
Functions on specific pages of GEO, PubMed, and DataMed. It has two functions: (1) to create downloadable citations for GEO data and PubMed articles and (2) to highlight the most important sentences in PubMed abstracts in a graded manner (based on TextRank algorithm).
Drug Pair Seeker
Predict and Prioritize Pairs of Drugs
A Java program that attempts to predict and prioritize pairs of drugs using the Connectivity Map dataset. Users can enter lists of up and down differentially expressed genes from their experiments to receive a ranked list of drug combinations that are predicted to either reverse or augment the gene expression state of the cells or tissue of interest using a simple formula.
PMID: 23559582
Network2Canvas
Perform and Visualize Gene-set and Drug-set Enrichment Analyses
A web application that provides an alternative way to view networks. N2C visualizes networks by placing nodes on a square toroidal canvas. The network nodes are clustered on the canvas using simulated annealing to maximize local connections where a node’s brightness is made proportional to its local fitness. The interactive canvas is implemented in HyperText Markup Language (HTML)5 with the JavaScript library Data-Driven Documents (D3). We applied N2C to visualize 30 canvases made from human and mouse gene-set libraries and 6 canvases made from the Food and Drug Administration (FDA)-approved drug-set libraries.
PMID: 23749960
ChEA
ChIP-X Enrichment Analysis
Database contains manually extracted datasets of transcription-factor/target-gene interactions from over 100 experiments such as ChIP-chip, ChIP-seq, ChIP-PET applied to mammalian cells. We use the database to analyze mRNA expression data where we perform gene-list enrichment analysis as the prior biological knowledge gene-list library. The system is delivered as web-based interactive software. With this software users can input lists of mammalian genes for which the program computes over-representation of transcription factor targets from the ChEA database.
PMID: 20709693
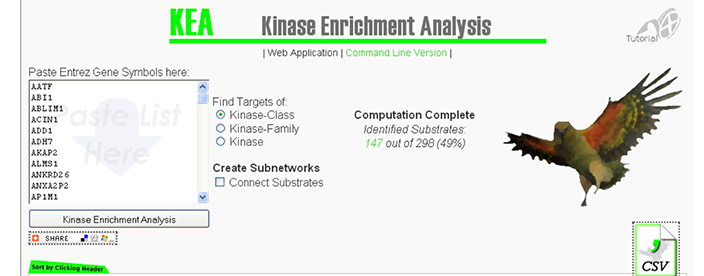
KEA
Kinase Enrichment Analysis
A web-based tool with an underlying database providing users with the ability to link lists of mammalian proteins/genes with the kinases that phosphorylate them. The system draws from several available kinase-substrate databases to compute kinase enrichment probability based on the distribution of kinase-substrate proportions in the background kinase-substrate database compared with kinases found to be associated with an input list of genes/proteins.
PMID: 19176546
GATE
Grid Analysis of Time-series Expression
A computational software platform for integrated visualization and analysis of expression time-series. Given a high-dimensional time-series dataset, GATE employs a clustering algorithm that creates movies of expression dynamics by assigning individual genes/proteins to hexagons on a hexagonal array and dynamically coloring each hexagon according to the expression level of the molecular species with which it is associated. Additionally, in order to infer potential regulatory control mechanisms from patterns of time-series correlations, GATE allows interactive interrogation of the movies with a wide variety of background knowledge datasets.
PMID: 19892805
Genes2FANs
Utilizes FANs and a PPI Network to Build Subnetworks that Connect Lists of Human and Mouse Genes
A web-based tool and a database that utilizes 14 carefully constructed functional association networks (FANs) and a large-scale protein-protein interaction (PPI) network to build subnetworks that connect input lists of human and mouse genes. The FANs are created from mammalian gene set libraries where mouse genes are converted to their human orthologs. The tool takes as input a list of human or mouse Entrez gene symbols to produce a subnetwork and a ranked list of intermediate genes that are used to connect the query input list. In addition, users can enter any PubMed search term and then the system automatically converts the returned results to gene lists using GeneRIF. This gene list is then used as input to generate a subnetwork from the user’s PubMed query.
PMID: 22748121
Sets2Networks
Network Inference from Repeated Observations of Sets
A general method for network inference from repeated observations of sets of related entities. Given experimental observations of sets of related entities, S2N infers the underlying network of binary interactions between these entities by generating an ensemble of networks consistent with the data; the frequency of occurrence of a given interaction throughout this ensemble is interpreted as the probability that the interaction is present in the underlying real network.
PMID: 22824380
Expression2Kinases
Gene Expression Data Analysis
A method to identify upstream regulators likely responsible for observed patterns in genome-wide gene expression. By integrating ChIP-seq/chip and position-weight-matrices (PWMs) data, protein-protein interactions, and kinase-substrate phosphorylation reactions, X2K can better identify regulatory mechanisms upstream of genome-wide differences in gene expression. X2K first infers the most likely transcription factors that regulate the differences in gene expression, then uses protein-protein interactions to connect the identified transcription factors using additional proteins for building transcriptional regulatory subnetworks centered on these factors, and finally uses kinase-substrate protein phosphorylation reactions, to identify and rank candidate protein-kinases that most likely regulate the formation of the identified transcriptional complexes.
PMID: 22080467
Lists2Networks
Integrated Analysis of Gene/Protein Lists
A web-based system that allows users to upload and analyze lists of mammalian gene-sets in a client-server software application. Within their workspace users can examine the overlap among the lists they upload, manipulate lists with different set operations, expand lists using existing mammalian networks of protein-protein, co-expression correlations, or background knowledge annotation correlations, and apply simple gene-set enrichment analyses on many gene lists at once against a plethora of prior knowledge datasets.
PMID: 20152038
Genes2Networks
Tool for Creating Subnetworks from Lists of Mammalian Genes or Proteins
A software tool that can be used to place lists of mammalian genes in the context of background mammalian signalome and interactome networks. The input to the program is a list of human Entrez Gene gene symbols and background networks in SIG format, while the output includes: (a) all identified interactions for the genes/proteins, (b) a subnetwork connecting the genes/proteins using intermediate components that are used to connect the genes, (c) ranking of the specificity of intermediate components to interact with the list of genes/proteins.
PMID: 17916244
Flash-based Network Viewer
Network Visualization
Visualization of small to moderately sized biological networks and pathways. FVN can also be used to embed pathways inside PDF files for the communication of pathways in soft publication materials.
PMID: 21349871
Genes2WordCloud
Identify Biological Themes from Gene Lists
A word-cloud generator and a word-cloud viewer that is based on WordCram implemented using Java, Processing, AJAX, mySQL, and PHP. Text is fetched from several sources and then processed to extract the most relevant terms with their computed weights based on word frequency.
PMID: 21995939
Sig2BioPAX
Tool for Converting Flat Files to BioPAX
A command-line Java program that can be used to convert structured text files describing molecular interactions into the BioPAX Level 3 standard format.
PMID: 21418653
SNAVI
Desktop Application for Analysis and Visualization of Large-Scale Cell Signaling Networks
A Windows-based desktop application that implements standard network analysis methods to compute the clustering, connectivity distribution, and detection of network motifs, as well as provides means to visualize networks and network motifs. SNAVI is capable of generating linked web pages from network datasets loaded in text format. SNAVI can also create networks from lists of gene or protein names. SNAVI is a useful tool for analyzing, visualizing and sharing cell signaling data. SNAVI is open source free software.
PMID: 19154595
AJAX Viewer for Signaling Networks
Web-based Viewer of Interactive Cell signaling Networks
A visualization tool for viewing and sharing intracellular signaling, gene regulation and protein interaction networks. AVIS is implemented as an AJAX-enabled syndicated Google gadget. It allows any webpage to render an image from a text file representation of signaling, gene regulatory, or protein interaction networks.
PMID: 17855420
PubMed Alert Me!
PubMed SDI Software Application
A software utility that allows users to enter a list of PubMed queries. Once a list of queries is configured, the program runs either daily or weekly. It searches PubMed and if it finds new matching published papers, the program sends an e-mail notification with a list of links to the new articles.
PMID: 18402930
ESCAPE
Database for Integrating High-content Data Collected from Human and Mouse Embryonic Stem Cells
A mammalian embryonic stem cell (ESC)-specific database created by collecting and integrating data reporting results from various published studies that profiled human and mouse ESCs including: protein-DNA binding interactions extracted from ChIP-seq/chip experiments, gene regulatory interactions from loss/gain-of-function studies followed by genome-wide mRNA expression profiling, protein interactions from immunoprecipitation followed by mass-spectrometry proteomics, a list of potential pluripotency regulators from RNA interference screens, ESC-specific proteins and phosphoproteins with specified phosphosites from proteomics and phosphoproteomics studies, time-course genome-wide mRNA microarray datasets from differentiating mouse ESCs, and histone modification status from genome-wide studies.
PMID: 23794736
Neuronal Signalome
Model Representing Signaling Pathways and Cellular Machines in the Hippocampal CA1 Neuron
Consists of cell signaling interactions extracted from literature describing components and interactions in mammalian neurons. This network integrates cell signaling pathways specific to mammalian neurons.
PMID: 16099987
Presynaptome
PRE Literature-based Protein-Protein Interaction Network
Consists of literature-based protein-protein interactions extracted from low-throughput experimental studies reporting interactions in mammalian presynaptic nerve terminals.
PMID: 19562802